15 Oct Dothiorella
Dothiorella Sacc., Michelia 2(6): 5 (1880)
Dothiorella was proposed by Saccardo to accommodate D. pyrenophora (Hyde et al. 2014). Members of this genus are pathogens, endophytes and saprobes (Phillips et al. 2013; Dissanayake et al. 2016). Taxonomy of this genus has been in a state of flux for decades (Phillips et al. 2013). Sivanesan (1984) treated D. pyrenophora as a synonym of Dothichiza sorbi (asexual morph of Dothiora pyrenophora). However, Sivanesan (1984) was referring to Dothiorella pyrenophora Sacc. (1884) which, according to Sutton (1977), is a later homonym of Dothiorella pyrenophora Sacc. (1880). Crous and Palm (1999) studied the holotype of D. pyrenophora and considered it a synonym of Diplodia. However, Phillips et al. (2005) based on both morphological and molecular data revived the genus Dothiorella for species in which the conidia become brown and 1-septate while attached to the conidiogenous cells.
Classification – Dothideomycetes, incertae sedis, Botryosphaeriales, Botryosphaeriaceae
Type species – Dothiorella pyrenophora Berk. ex Sacc., Michelia 2 (6): 5 (1880) (1909)
Distribution – Worldwide
Disease symptoms – Diebacks, cankers, fruit rots.
Hosts – Plurivorous on woody hosts.
Morphological based identification and diversity
Species of this genus were mostly described based on host association, which has led to the introduction of many species names and currently there are 393 epithets in Index Fungorum (2019). Slippers et al. (2013) suggested that host association cannot be considered as an important factor in species delimitation, many names are likely to be synonyms. Phillips et al. (2008) introduced a new genus Spencermartinsia to accommodate dothiorella-like species with apiculate ascospores. However, Yang et al. (2017) based on six-gene phylogeny and a broad taxon sampling considered that Spencermartinsia should be treated as a synonym of Dothiorella. Phillips et al. (2013) listed all cultures available for this genus and provided a phylogenetic tree and a key to the species. In that study 13 species, names and 16 unnamed lineages were listed. Hyde et al. (2014), Dissanayake et al. (2016) and Yang et al. (2017) provided updates for the genus. Hyde et al. (2014) accepted 19 species and Dissanayake et al. (2016) accepted 30 species in the genus. After making Spencermartinsia a synonym of Dothiorella Yang et al. (2017) accepted 36 species in this genus.
Phillips et al. (2013) differentiated 13 Dothiorella species on the basis of conidiomata and conidial dimensions. However, the dimensions of these characters overlap between species. Therefore, using morphology alone without molecular data is not suitable to define species.
Molecular based identification and diversity
Recent studies have re-evaluated this genus based on multi-gene phylogeny of ITS, TUB2 and tef1 sequence data. We reconstruct the phylogeny of Dothiorella based on analyses of a combined ITS and tef1 sequence data (Table, Fig). The phylogenetic tree is updated with recently introduced Dothiorella species and corresponds to previous studies (Dissanayake et al. 2016; Yang et al. 2017; Hyde et al. 2019; Phookamsak et al. 2019). In the analyses, it appears that several species are synonyms, such as D. parva/D. guttulata and D. rhamni/D. eriobotryae and possibly others. Therefore, a thorough revision of the genus is recommended to clarify the status of these dubious species.
Recommended genetic markers (genus level) – SSU and LSU
Recommended genetic markers (species level) – ITS and tef1
Accepted number of species: Currently, 393 species names are listed for Dothiorella in Index Fungorum (2019). Cultures and DNA sequences are available for 46 species, therefore 46 species are currently accepted in Dothiorella.
.
References: Phillips et al. 2013; Dissanayake et al. 2016 (morphology, phylogeny, distribution, hosts), Yang et al. 2017 (morphology and phylogeny)
Table Details of the Dothiorella isolates used in the phylogenetic analyses. Ex-type (ex-epitype) strains are in bold and marked with an asterisk* and voucher strains are in bold.
| Species | Isolate/Voucher No | ITS | tef1 |
| Dothiorella acacicola | CBS 141295 | KX228269 | KX228376 |
| D. acericola | KUMCC 18-0137* | MK359449 | MK361182 |
| D. alpina | CGMCC 3.18001* | KX499645 | KX499651 |
| D. americana | CBS 128309* | HQ288218 | HQ288262 |
| D. brevicollis | CBS 130411* | JQ239403 | JQ239390 |
| D. californica | CBS 141587* | KX357188 | KX357211 |
| D. capri-amissi | CMW 25403* | EU101323 | EU101368 |
| D. casuarinae | CBS 120688* | DQ846773 | DQ875331 |
| D. citricola | ICMP16828* | EU673323 | EU673290 |
| D. dulcispinae | CBS 130413* | JQ239400 | JQ239387 |
| D. eriobotryae | CBS 140852* | KT240287 | KT240262 |
| D. guttulataa | MFLUCC 17-0242 | KY797637 | – |
| D. iberica | CBS 115041* | AY573202 | AY573222 |
| D. iranica | IRAN1587C* | KC898231 | KC898214 |
| D. italica | MFLUCC 17-0951 | MG828897 | MG829267 |
| D. juglandis | CBS 188.87 | EU673316 | EU673283 |
| D. lampangensis | MFLUCC 18-0232 | MK347758 | MK340869 |
| D. longicollis | CBS 122068* | EU144054 | EU144069 |
| D. magnoliae | CFCC 51563 | KY111247 | KY213686 |
| D. mangifericola | IRAN1584C* | KC898221 | KC898204 |
| D. moneti | MUCC505* | EF591920 | EF591971 |
| D. neclivorem | DAR80992* | KJ573643 | KJ573640 |
| D. oblonga | CMW 25407* | EU101300 | EU101345 |
| D. omnivora | CBS 140349* | KP205497 | KP205470 |
| D. parva | IRAN1579C* | KC898234 | KC898217 |
| D. plurivora | IRAN1557C* | KC898225 | KC898208 |
| D. pretoriensis | CBS 130404* | JQ239405 | JQ239392 |
| D. prunicola | CBS 124723* | EU673313 | EU673280 |
| D. rhamni | MFLUCC 14-0902* | KU246381 | – |
| D. rosulata | CBS 121760* | EU101290 | EU101335 |
| D. santali | MUCC 509* | EF591924 | EF591975 |
| D. sarmentorum | IMI63581b* | AY573212 | AY573235 |
| D. sempervirentis | IRAN1583C* | KC898236 | KC898219 |
| D. striata | ICMP16824* | EU673320 | EU673287 |
| D. styphnolobii | JZB3150013* | MH880849 | MK069594 |
| D. symphoricarposicola | MFULCC 13-0497* | KJ742378 | KJ742381 |
| D. tectonae | MFLUCC12-0382* | KM396899 | KM409637 |
| D. thailandica | CBS 133991* | JX646796 | JX646861 |
| D. thripsita | BRIP 51876* | FJ824738 | KJ573639 |
| D. ulmacea | CBS 138855* | KR611881 | KR611910 |
| D. uruguayensis | CBS 124908* | EU080923 | EU863180 |
| D. vidmadera | DAR78992* | EU768874 | EU768881 |
| D. vinea-gemmae | DAR81012* | KJ573644 | KJ573641 |
| D. viticola | CBS 117009* | AY905554 | AY905559 |
| D. westrale | DAR80529* | HM009376 | HM800511 |
| D. yunnana | CGMCC 3.17999* | KX499643 | KX499649 |
a the tef1 sequence of D. guttulata in GenBank is incorrect, therefore was not included in the analyses.
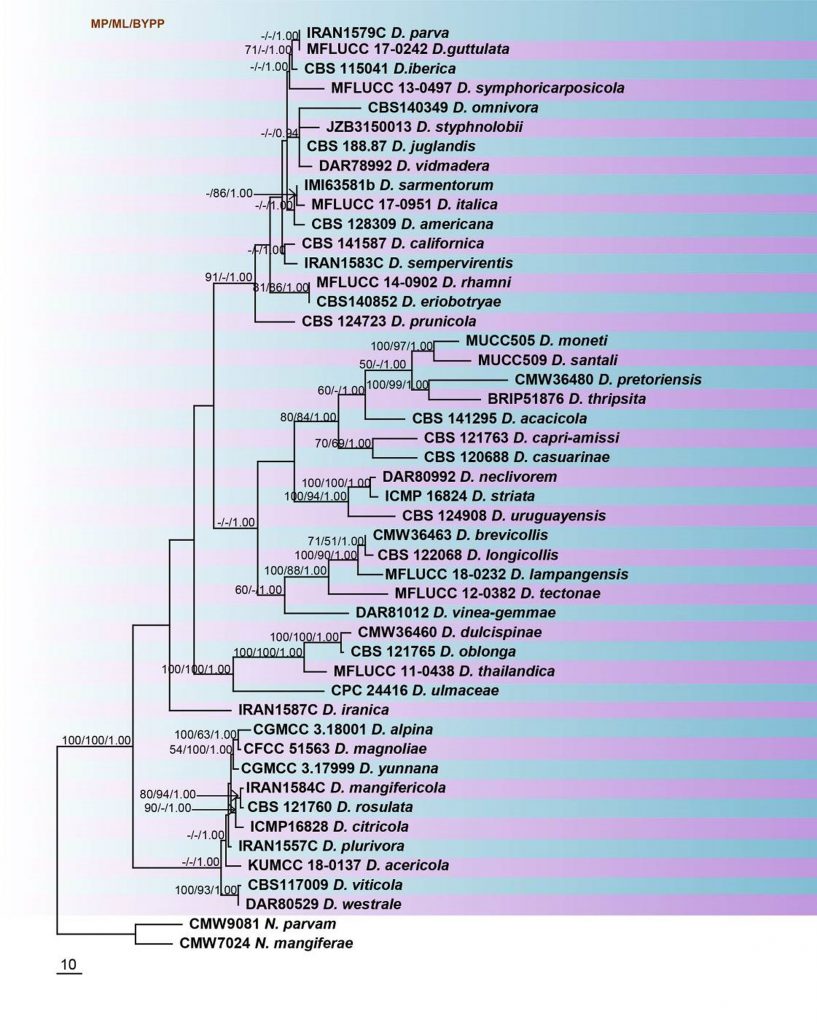
Fig. Phylogenetic tree generated by maximum parsimony analysis of combined ITS and tef1 sequence data of Dothiorella species. Related sequences were obtained from GenBank. Forty-eight strains are included in the analyses, which comprised 873 characters including gaps. The tree was rooted with Neofusicoccum parvum (CMW9081) and N. mangiferae (CMW7024). The maximum parsimonious dataset consisted of 572 constant, 204 parsimony-informative and 97 parsimony-uninformative characters. The parsimony analysis of the data matrix resulted in the maximum of ten equally most parsimonious trees with a length of 891 steps (CI = 0.532, RI 0.738, RC = 0.393, HI = 0.468) in the first tree. MP and ML bootstrap values ≥50% and Bayesian posterior probabilities ≥0.90 are shown respectively near the nodes. The scale bar indicates 10 changes per site. Ex-type strains are in bold.

No Comments